Jun 09, 2022
Given two matrices \(A \in \mathcal{R}^{M\times S}\) and \(B \in \mathcal{R}^{S \times N}\) it is helpful to be able to reorder their product \(A B \in \mathcal{R}^{M\times N}\) in order to compute Jacobians for optimization. For this it is common to define the \(\mbox{vec}(.)\) operator which flattens a matrix into a vector by concatenating columns:
\begin{equation*}
\newcommand{\vec}[1]{\mbox{vec}{\left(#1\right)}}
\vec{A} = \begin{bmatrix} A_{11} \\ \vdots \\ A_{M1} \\ \vdots \\ A_{1S} \\ \vdots \\ A_{MS}\end{bmatrix}
\end{equation*}
Using this operator, the (flattened) product \(AB\) can be written as one of:
\begin{align*}
\vec{AB} &=& (B^T \otimes I_A)\vec{A} \\
&=& (I_B \otimes A)\vec{B}
\end{align*}
where \(\otimes\) is the Kronecker Product and \(I_A\) is an identity matrix with the same number of rows as \(A\) and \(I_B\) is an idenity matrix with the same number of columns as \(B\).
From the two options for expressing the product, it's clear that \((B^T \otimes I_A)\) is the (vectorized) Jacobian of \(AB\) with respect to \(A\) and \((I_B \otimes A)\) is the corresponding vectorized Jacobian with respect to \(B\).
That's great but if our linear algebra library happens to represent matrices in row-major order it will be necessary to sprinkle a large number of transposes in to get things in the right order to apply these formulas. That will be tedious, error-prone and inefficient. Fortunately, equivalent formulas are available for row-major ordering by defining a \(\mbox{ver}(.)\) operator that flattens a matrix by concatenating rows to form a column vector:
\begin{equation*}
\newcommand{\ver}[1]{\mbox{ver}{(#1)}}
\ver{A} = \begin{bmatrix} A_{11} \\ \vdots \\ A_{1S} \\ \vdots \\ A_{M1} \\ \vdots \\ A_{MS} \end{bmatrix}
\end{equation*}
Given this it is fairly quick to work out that the equivalent to the first two formulas using the \(\ver{.}\) operator are:
\begin{align*}
\ver{AB} &=& (I_A \otimes B^T)\ver{A} \\
&=& ( A \otimes I_B)\ver{B}
\end{align*}
Using the same logic it can be concluded that \((I_A \otimes B^T)\) is the vectorized Jacobian of \(AB\) with respect to \(A\) and \((A \otimes I_B)\) is the corresponding Jacobian with respect to \(B\).
It's also worth noting that these are simplifications of a more general case for the product \(AXB\) that brings \(X\) to the right of the expression. The corresponding \(\vec{.}\) and \(\ver{.}\) versions are:
\begin{align*}
\vec{AXB} &=& (B^T \otimes A) \vec{X} \\
\ver{AXB} &=& (A \otimes B^T) \ver{X}
\end{align*}
which makes it clear where the identity matrices \(I_A\) and \(I_B\) in the previous expressions originate.
All these can be demonstrated by the following python code sample:
from typing import Tuple
import numpy as np
def vec( M: np.ndarray ):
'''Flatten argument by columns'''
return M.flatten('F')
def ivec( v: np.ndarray, shape: Tuple[int,int] ):
'''Inverse of vec'''
return np.reshape( v, shape, 'F' )
def ver( M: np.ndarray ):
'''Flatten argument by rows'''
return M.flatten('C')
def iver( v: np.ndarray, shape: Tuple[int,int] ):
'''Inverse of ver'''
return np.reshape( v, shape, 'C' )
def test_vec_ivec():
A = np.random.standard_normal((3,4))
assert np.allclose( A, ivec( vec( A ), A.shape ) )
def test_ver_iver():
A = np.random.standard_normal((3,4))
assert np.allclose( A, iver( ver(A), A.shape ) )
def test_products():
A = np.random.standard_normal((3,4))
X = np.random.standard_normal((4,4))
B = np.random.standard_normal((4,5))
AB1 = A@B
Ia = np.eye( A.shape[0] )
Ib = np.eye( B.shape[1] )
AB2a = ivec( np.kron( B.T, Ia )@vec(A), (A.shape[0],B.shape[1]) )
AB2b = ivec( np.kron( Ib, A )@vec(B), (A.shape[0],B.shape[1]) )
assert np.allclose( AB1, AB2a )
assert np.allclose( AB1, AB2b )
AB3a = iver( np.kron( A, Ib )@ver(B), (A.shape[0], B.shape[1]) )
AB3b = iver( np.kron( Ia, B.T )@ver(A), (A.shape[0], B.shape[1]) )
assert np.allclose( AB1, AB3a )
assert np.allclose( AB1, AB3b )
AXB1 = A@X@B
AXB2 = ivec( np.kron(B.T,A)@vec(X), (A.shape[0],B.shape[1]) )
AXB3 = iver( np.kron(A,B.T)@ver(X), (A.shape[0],B.shape[1]) )
assert np.allclose( AXB1, AXB2 )
assert np.allclose( AXB1, AXB3 )
if __name__ == '__main__':
test_vec_ivec()
test_ver_iver()
test_products()
Hopefully this is helpful to someone.
Dec 03, 2021
These are some helpful quaternion identities, all summarized from Kok et al., Using Inertial Sensors for Position and Orientation Estimation although I have put the scalar component last.
A quaternion \(q\) is represented as a 4-tuple, or an imaginary vector portion \(q_v=[q_x,q_y,q_z]^T\) and a scalar value \(q_s\) giving \(q = [q_v^T, q_s]^T\). The norm of a quaternion is \(\|q\|_2 = (q_v^T q_v + q_s^2)^{\frac{1}{2}}\), the conjugate of a quaternion is \(q^* = [-q_v^T, q_s]^T\) and the inverse of a quaternion is \(q^{-1} = \frac{q^*}{\|q\|_2^2}\).
Unit quaternions have \(\|q\|_2 = 1\), in which case \(q^* = q^{-1}\). Unit quaternions also represent spatial rotations/orientations of \(\theta\) radians around a unit axis \(a\) via \(q = [a^T \sin\left(\frac{\theta}{2}\right), \cos\left(\frac{\theta}{2}\right) ]^T\). Both \(q\) and \(-q = [-q_v, -q_s]^T\) represent the same orientation, although the one with positive scalar component takes the shortest rotation to achieve that orientation since \(\cos^{-1}(|x|) \leq \cos^{-1}(-|x|)\).
The multiplication of two quaternions is \(p \otimes q = [ (p_s q_v + q_s p_v + p_v \times q_v)^T, p_s q_s - p_v \times q_v ]^T\). The presence of the cross-product means that \(p\otimes q \neq q \otimes p\). When \(p, q\) are unit quaternions, \(p\otimes q\) is equivalent to chaining the two rotations. The rotation matrix corresponding to a unit quaternion is:
\begin{equation*}
R(q) = \begin{bmatrix} 1 - 2 q_y^2 - 2*q_z^2 & 2 q_x q_y - 2 q_z q_s & 2 q_x q_z + 2 q_y q_s \\
2 q_x q_y + 2 q_z q_s & 1 - 2 q_x^2 - 2 q_z^2 & 2 q_y q_z - 2 q_x q_s \\
2 q_x q_z - 2 q_y q_s & 2 q_y q_z + 2 q_x q_s & 1 - 2 q_x^2 - 2 q_y^2 \end{bmatrix}
\end{equation*}
or in python:
def quat2mat( q: np.ndarray ) -> np.ndarray:
assert q.ndim == 1 and q.size == 4
qx,qy,qx,qs = q
return np.ndarray((
( 1 - 2*qy**2 - 2*qz**2, 2*qx*qy - 2*qz*qs, 2*qx*qz + 2*qy*qs ),
( 2*qx*qy + 2*qz*qs, 1 - 2*qx**2 - 2*qz**2, 2*qy*qz - 2*qx*qs ),
( 2*qx*qz - 2*qy*qs, 2*qy*qz + 2*qx*qs, 1 - 2*qx**2 - 2*qy**2 )
))
The, usually non-unit, quaternion representation of a vector \(v^\wedge = [ v^T, 0]^T\). The vector can be rotated by the unit-quaternion \(q\) via \(R(q) v = (q\otimes v^\wedge \otimes q^*)_v\) where only the vector portion is retained.
The quaternion-quaternion product can be represented by matrix multiplication in either the same, or reversed order, i.e.:
\begin{align*}
\newcommand{\pd}[2]{\frac{\partial #1}{\partial #2}}
p \otimes q &= p^L q = q^R p \hspace{1cm} & \mbox{where} \\
p^L &= \begin{bmatrix} p_s I + \lfloor p_v \rfloor_\times & p_v \\ -p_v^T & p_s \end{bmatrix} & \\
q^R &= \begin{bmatrix} q_s I - \lfloor q_v \rfloor_\times & q_v \\ -q_v^T & q_s \end{bmatrix} &
\end{align*}
where \(\lfloor m \rfloor_\times\) is the matrix representation of the cross-product. This is extremely useful when differentiating quaternion expressions since \(p^L = \pd{}{q}(p\otimes q)\) and \(q^R = \pd{}{p}(p\otimes q)\).
A Rodrigues rotation vector \(\eta = \theta a\) where \(\|a\|_2 = 1\) can be mapped to a quaternion via the exponential map:
\begin{align*}
\exp_q(\eta) =& q = \begin{pmatrix} a \sin\left(\frac{\theta}{2}\right) \\ \cos\left(\frac{\theta}{2}\right) \end{pmatrix} \\
\log_q(q) =& \eta = 2 q_v
\end{align*}
Under the assumption that \(\theta << 1\), the following approximations can be made:
\begin{align*}
\exp_q(\eta) \approx& \begin{pmatrix} \frac{\eta}{2} \\ 1 \end{pmatrix} \\
\pd{\exp_q(\eta)}{\eta} \approx& \begin{pmatrix} \frac{1}{2} I_{3\times3} \\ 0 \end{pmatrix} \\
\pd{\log_q(q)}{q} \approx& \begin{pmatrix} 2 I_{3\times3} & 0_{3\times1} \end{pmatrix}
\end{align*}
The above are also very helpful in manifold optimization over quaternion states where the Newton update is generally small which results in linear Jacobians with respect to the update variables.
Nov 28, 2021
I find this process easier to reason through by writing out a small 3-column version explicitly:
\begin{equation*}
\begin{bmatrix} A_{*0} & A_{*1} & A_{*2} \end{bmatrix} =
\begin{bmatrix} Q_{*0} & Q_{*1} & Q_{*2} \end{bmatrix} \begin{bmatrix}
R_{00} & R_{01} & R_{02} \\
& R_{11} & R_{12} \\
& & R_{22}
\end{bmatrix}
\end{equation*}
which gives:
\begin{align*}
&A_{*0} = Q_{*0} R_{00} \\
&A_{*1} = Q_{*0} R_{01} + Q_{*1} R_{11} \\
&A_{*2} = Q_{*0} R_{02} + Q_{*1} R_{12} + Q_{*2} R_{22}
\end{align*}
From this and the orthonormal properties of \(Q\), the meaning of the \(R_{ji}\) values are clear, the projection of the \(i\)'th column of \(A\) onto the \(j\)'th column of \(Q\). These allows fixing the values of \(R_{ji}\) and norms of \(Q_{*i}\):
- \(Q_{*i}^T Q_{*j} = 0, \forall i \neq j\): columns of \(Q\) are orthogonal so each \(R_{j*}\) value must account for entire parallel component of \(A_{*j}\) onto each column of \(Q\).
- \(\|Q_{*i}\|_2 = 1, \forall i\): constrains \(\|Q_{*i}\|\) and corresponding \(R_{ji}\) values.
Therefore:
- \(R_{00} = \|A_{*0}\|_2\) and \(Q_{*0} = A_{*0}/R_{00}\)
- \(R_{01} = Q_{*0}^T A_{*1}\), \(s_1 = A_{*1} - R_{01} Q_{*0}\), \(R_{11} = \|s_1\|_2\) \(Q_{*1} = s_1/R_{11}\)
- \(R_{02} = Q_{*0}^T A_{*2},~R_{12} = Q_{*1}^T A_{*2}, s_2 = A_{*2} - R_{02} Q_{*0} - R_{12} Q_{*1}, R_{22} = \| s_2 \|_2, Q_{*2} = s_2/R_{22}\)
or in general:
- \(R_{ji} = Q_{*j}^T A_{*i}, \forall j < i\)
- \(s_{i} = A_{*i} - \sum_j R_{ji} Q_{*j}\)
- \(R_{ii} = \| s_{i} \|_2, Q_{*i} = s_i/R_{ii}\)
and in python:
def qr_gs( A, inplace=True ):
'''Decompose A into Q*R with Q orthonormal and R upper triangular using classical Gram-Schmidt (unstable)'''
A = A if inplace else A.copy()
R = np.zeros((A.shape[1],A.shape[1]))
for i in range( A.shape[1] ):
for j in range(i):
R[j,i] = np.sum(A[:,j]*A[:,i])
A[:,i] -= R[j,i]*A[:,j]
R[i,i] = np.linalg.norm(A[:,i])
A[:,i] /= R[i,i]
return A,R
The form above generates \(Q\) column by column but has stability issues due to the use of classical Gram-Schmidt. It can be improved by replacing classical Gram-Schmidt with modified Gram-Schmidt:
def qr_mgs( A, inplace=True ):
'''Decompose A into Q*R with Q orthonormal and R upper triangular using modified Gram-Schmidt
Assumes columns of A are linearly independent.
'''
A = A if inplace else A.copy()
R = np.zeros((A.shape[1],A.shape[1]))
for i in range( A.shape[1] ):
R[i,i] = np.linalg.norm(A[:,i])
A[:,i] /= R[i,i]
for j in range(i+1,A.shape[1]):
R[i,j] = np.dot(A[:,j],A[:,i])
A[:,j] -= R[i,j]*A[:,i]
return A,R
The inner loop of this can be replaced entirely with vectorization in python but the above form gives a reasonable starting point for porting to another language like C or C++. These implementations should also check for \(R_{ii} = 0\) which indicate a rank deficient system.
Jun 14, 2021
Given a linear model \(y_i = x_i^T \beta\), we collect \(N\) observations \(\tilde{y_i} = y_i + \epsilon_i\) where \(\epsilon_i \sim \mathcal{N}(0,\sigma^2)\). The likelihood of any one of our observations \(\tilde{y_i}\) given current estimates of our parameters \(\beta\) is then:
\begin{equation*}
\newcommand{\exp}[1]{{\mbox{exp}\left( #1 \right)}}
\newcommand{\log}[1]{{\mbox{log}\left( #1 \right)}}
\DeclareMathOperator*{\argmax}{argmax}
\DeclareMathOperator*{\argmin}{argmin}
P(\tilde{y}_i | \beta) = \frac{1}{\sqrt{2\pi\sigma^2}} \exp{-\frac{1}{2}\left(\frac{e_i}{\sigma}\right)^2} = \frac{1}{\sqrt{2\pi\sigma^2}} \exp{-\frac{1}{2}\left(\frac{\tilde{y_i}-x_i^T \beta}{\sigma}\right)^2}
\end{equation*}
and the likelihood of all of them occuring is:
\begin{equation*}
P(\tilde{y} | \beta) = \prod_{i=1}^N P(\tilde{y}_i | \beta )
\end{equation*}
The parameters most likely to explain the observations, given only the observations and definition of the model, are those that maximize this likelihood. Equivalently, we can find the parameters that minimize the negative log-likelihood since the two have the same optima:
\begin{align*}
\beta^* &=& \argmax_\beta P(\tilde{y} | \beta) \\
&=& \argmin_\beta - \ln( P(\tilde{y} | \beta ) \\
&=& \argmin_\beta - N\log{\frac{1}{\sqrt{2\pi\sigma^2}}} - \log{ \prod_{i=1}^N P(\tilde{y}_i | \beta ) }
\end{align*}
The benefit of this is that the terms decouple in the case of normally distributed residuals. Ditching constant factors and collecting the product terms into the exponent the above becomes:
\begin{align*}
\beta^* &=& \argmin_\beta -\ln\left( \exp{-\frac{1}{2}\sum_{i=1}^N \left(\frac{\tilde{y}_i - M_i^T \beta}{\sigma}\right)^2} \right) \\
&=& \argmin_\beta \frac{1}{2\sigma^2} \sum_{i=1}^N \left( \tilde{y}_i - M_i^T \beta \right)^2
\end{align*}
which gives the least-squares version.
Least Squares & MAP
We can also add a prior for the parameters \(\beta\), e.g. \(\beta_i \sim \mathcal{N}(0,\sigma_\beta)\). In this case, each component of \(\beta\) is independent. The MAP estimate for \(\beta\) is then:
\begin{align*}
\beta^* &=& \argmax_\beta P(\tilde{y} | \beta) P(\beta) \\
&=& \argmax_\beta \prod_i P(\tilde{y}_i | \beta) \prod_j P(\beta_j) \\
&=& \argmax_\beta \prod_i \frac{1}{\sqrt{2\pi\sigma^2}}\exp{ -\frac{1}{2}\left(\frac{e_i}{\sigma}\right)^2} \prod_j \frac{1}{\sqrt{2\pi\sigma_\beta^2}}\exp{ -\frac{1}{2}\left(\frac{\beta_j}{\sigma_\beta}\right)^2 }
\end{align*}
The same negative log trick can then be used to convert products to sums, also ditching constants, to get:
\begin{align*}
\beta^* &=& \argmin \frac{1}{2\sigma^2} \sum_i e_i^2 + \frac{1}{2\sigma_\beta^2} \sum_j \beta_j^2 \\
&=& \argmin \frac{1}{2\sigma^2} \sum_i \left( \tilde{y}_i - x_i^T \beta \right)^2 + \frac{1}{2\sigma_\beta^2} \sum_j \beta_j^2
\end{align*}
Using different priors will of course give different forms for the second term, e.g. choosing components of \(\beta\) to follow a Laplace distribution \(\beta_j \sim \frac{\lambda}{2} \exp{-\lambda |\beta_j|}\), gives:
\begin{equation*}
\beta^* = \argmin \frac{1}{2\sigma^2} \sum_i \left( \tilde{y}_i - x_i^T \beta \right)^2 + \lambda \sum_j |\beta_j|
\end{equation*}
Dec 13, 2020
The essential matrix relates normalized coordinates in one view with those in another view of the same scene and is widely used in stereo reconstruction algorithms. It is a 3x3 matrix with rank 2, having only 8 degrees of freedom.
Given \(p\) as a normalized point in image P and \(q\) as the corresponding normalied point in another image Q of the same scene, the essential matrix provides the constraint:
\begin{equation*}
q^T E p = 0
\end{equation*}
It's important that \(p\) and \(q\) be normalized homogeneous points, i.e. the following where \([u_p,v_p]\) are the pixel coordinates of point p:
\begin{equation*}
p = K^{-1} \begin{bmatrix} u_p \\ v_p \\ 1 \end{bmatrix}
\end{equation*}
By fixing the extrinsics of image P as \(R_P = I\) and \(t_P = [0,0,0]^T\) the essential matrix relating P and Q can defined by the relationship:
\begin{equation*}
E = t_\times R
\end{equation*}
where \(R\) and \(t\) are the relative transforms of camera Q with respect to camera P and \(t_\times\) is the matrix representation of the the cross-product:
\begin{equation*}
t_\times = \begin{bmatrix} 0 & -t_z & t_y \\ t_z & 0 & -t_x \\ -t_y & t_x & 0 \end{bmatrix}
\end{equation*}
Nov 04, 2019
These are collections of functions that I end up re-implementing frequently.
from typing import *
import numpy as np
def closest_rotation( M: np.ndarray ):
U,s,Vt = np.linalg.svd( M[:3,:3] )
if np.linalg.det(U@Vt) < 0:
Vt[2] *= -1.0
return U@Vt
def mat2quat( M: np.ndarray ):
if np.abs(np.linalg.det(M[:3,:3])-1.0) > 1e-5:
raise ValueError('Matrix determinant is not 1')
if np.abs( np.linalg.norm( M[:3,:3].T@M[:3,:3] - np.eye(3)) ) > 1e-5:
raise ValueError('Matrix is not orthogonal')
w = np.sqrt( 1.0 + M[0,0]+M[1,1]+M[2,2])/2.0
x = (M[2,1]-M[1,2])/(4*w)
y = (M[0,2]-M[2,0])/(4*w)
z = (M[1,0]-M[0,1])/(4*w)
return np.array((x,y,z,w),float)
def quat2mat( q: np.ndarray ):
qx,qy,qz,qw = q/np.linalg.norm(q)
return np.array((
(1 - 2*qy*qy - 2*qz*qz, 2*qx*qy - 2*qz*qw, 2*qx*qz + 2*qy*qw),
( 2*qx*qy + 2*qz*qw, 1 - 2*qx*qx - 2*qz*qz, 2*qy*qz - 2*qx*qw),
( 2*qx*qz - 2*qy*qw, 2*qy*qz + 2*qx*qw, 1 - 2*qx*qx - 2*qy*qy),
))
if __name__ == '__main__':
for i in range( 1000 ):
q = np.random.standard_normal(4)
q /= np.linalg.norm(q)
A = quat2mat( q )
s = mat2quat( A )
if np.linalg.norm( s-q ) > 1e-6 and np.linalg.norm( s+q ) > 1e-6:
raise ValueError('uh oh.')
R = closest_rotation( A+np.random.standard_normal((3,3))*1e-6 )
if np.linalg.norm(A-R) > 1e-5:
print( np.linalg.norm( A-R ) )
raise ValueError('uh oh.')
Nov 03, 2019
Python is a pretty nice language for prototyping and (with numpy and scipy) can be quite fast for numerics but is not great for algorithms with frequent tight loops. One place I run into this frequently is in graphics, particularly applications involving meshes.
Not much can be done if you need to frequently modify mesh structure. Luckily, many algorithms operate on meshes with fixed structure. In this case, expressing algorithms in terms of block operations can be much faster. Actually implementing methods in this way can be tricky though.
To gain more experience in this, I decided to try implementing the As-Rigid-As-Possible Surface Modeling mesh deformation algorithm in Python. This involves solving moderately sized Laplace systems where the right-hand-side involves some non-trivial computation (per-vertex SVDs embedded within evaluation of the Laplace operator). For details on the algorithm, see the link.
As it turns out, the entire implementation ended up being just 99 lines and solves 1600 vertex problems in just under 20ms (more on this later). Full code is available.
Building the Laplace operator
ARAP uses cotangent weights for the Laplace operator in order to be relatively independent of mesh grading. These weights are functions of the angles opposite the edges emanating from each vertex. Building this operator using loops is slow but can be sped up considerably by realizing that every triangle has exactly 3 vertices, 3 angles and 3 edges. Operations can be evaluated on triangles and accumulated to edges.
Here's the Python code for a cotangent Laplace operator:
from typing import *
import numpy as np
import scipy.sparse as sparse
import scipy.sparse.linalg as spla
def build_cotan_laplacian( points: np.ndarray, tris: np.ndarray ):
a,b,c = (tris[:,0],tris[:,1],tris[:,2])
A = np.take( points, a, axis=1 )
B = np.take( points, b, axis=1 )
C = np.take( points, c, axis=1 )
eab,ebc,eca = (B-A, C-B, A-C)
eab = eab/np.linalg.norm(eab,axis=0)[None,:]
ebc = ebc/np.linalg.norm(ebc,axis=0)[None,:]
eca = eca/np.linalg.norm(eca,axis=0)[None,:]
alpha = np.arccos( -np.sum(eca*eab,axis=0) )
beta = np.arccos( -np.sum(eab*ebc,axis=0) )
gamma = np.arccos( -np.sum(ebc*eca,axis=0) )
wab,wbc,wca = ( 1.0/np.tan(gamma), 1.0/np.tan(alpha), 1.0/np.tan(beta) )
rows = np.concatenate(( a, b, a, b, b, c, b, c, c, a, c, a ), axis=0 )
cols = np.concatenate(( a, b, b, a, b, c, c, b, c, a, a, c ), axis=0 )
vals = np.concatenate(( wab, wab,-wab,-wab, wbc, wbc,-wbc,-wbc, wca, wca,-wca, -wca), axis=0 )
L = sparse.coo_matrix((vals,(rows,cols)),shape=(points.shape[1],points.shape[1]), dtype=float).tocsc()
return L
Inputs are float \(3 \times N_{verts}\) and integer \(N_{tris} \times 3\) arrays for vertices and triangles respectively. The code works by making arrays containing vertex indices and positions for all triangles, then computes and normalizes edges and finally computes angles and cotangents. Once the cotangents are available, initialization arrays for a scipy.sparse.coo_matrix are constructed and the matrix built. The coo_matrix is intended for finite-element matrices so it accumulates rather than overwrites duplicate values. The code is definitely faster and most likely fewer lines than a loop based implementation.
Evaluating the Right-Hand-side
The Laplace operator is just a warm-up exercise compared to the right-hand-side computation. This involves:
- For every vertex, collecting the edge-vectors in its one-ring in both original and deformed coordinates
- Computing the rotation matrices that optimally aligns these neighborhoods, weighted by the Laplacian edge weights, per-vertex. This involves a SVD per-vertex.
- Rotating the undeformed neighborhoods by the per-vertex rotation and summing the contributions.
In order to vectorize this, I constructed neighbor and weight arrays for the Laplace operator. These store the neighboring vertex indices and the corresponding (positive) weights from the Laplace operator. A catch is that vertices have different numbers of neighbors. To address this I use fixed sized arrays based on the maximum valence in the mesh. These are initialized with default values of the vertex ids and weights of zero so that unused vertices contribute nothing and do not affect matrix structure.
To compute the rotation matrices, I used some index magic to accumulate the weighted neighborhoods and rely on numpy's batch SVD operation to vectorize over all vertices. Finally, I used the same index magic to rotate the neighborhoods and accumulate the right-hand-side.
The code to accumulate the neighbor and weight arrays is:
def build_weights_and_adjacency( points: np.ndarray, tris: np.ndarray, L: Optional[sparse.csc_matrix]=None ):
L = L if L is not None else build_cotan_laplacian( points, tris )
n_pnts, n_nbrs = (points.shape[1], L.getnnz(axis=0).max()-1)
nbrs = np.ones((n_pnts,n_nbrs),dtype=int)*np.arange(n_pnts,dtype=int)[:,None]
wgts = np.zeros((n_pnts,n_nbrs),dtype=float)
for idx,col in enumerate(L):
msk = col.indices != idx
indices = col.indices[msk]
values = col.data[msk]
nbrs[idx,:len(indices)] = indices
wgts[idx,:len(indices)] = -values
Rather than break down all the other steps individually, here is the code for an ARAP class:
class ARAP:
def __init__( self, points: np.ndarray, tris: np.ndarray, anchors: List[int], anchor_weight: Optional[float]=10.0, L: Optional[sparse.csc_matrix]=None ):
self._pnts = points.copy()
self._tris = tris.copy()
self._nbrs, self._wgts, self._L = build_weights_and_adjacency( self._pnts, self._tris, L )
self._anchors = list(anchors)
self._anc_wgt = anchor_weight
E = sparse.dok_matrix((self.n_pnts,self.n_pnts),dtype=float)
for i in anchors:
E[i,i] = 1.0
E = E.tocsc()
self._solver = spla.factorized( ( self._L.T@self._L + self._anc_wgt*E.T@E).tocsc() )
@property
def n_pnts( self ):
return self._pnts.shape[1]
@property
def n_dims( self ):
return self._pnts.shape[0]
def __call__( self, anchors: Dict[int,Tuple[float,float,float]], num_iters: Optional[int]=4 ):
con_rhs = self._build_constraint_rhs(anchors)
R = np.array([np.eye(self.n_dims) for _ in range(self.n_pnts)])
def_points = self._solver( self._L.T@self._build_rhs(R) + self._anc_wgt*con_rhs )
for i in range(num_iters):
R = self._estimate_rotations( def_points.T )
def_points = self._solver( self._L.T@self._build_rhs(R) + self._anc_wgt*con_rhs )
return def_points.T
def _estimate_rotations( self, def_pnts: np.ndarray ):
tru_hood = (np.take( self._pnts, self._nbrs, axis=1 ).transpose((1,0,2)) - self._pnts.T[...,None])*self._wgts[:,None,:]
rot_hood = (np.take( def_pnts, self._nbrs, axis=1 ).transpose((1,0,2)) - def_pnts.T[...,None])
U,s,Vt = np.linalg.svd( rot_hood@tru_hood.transpose((0,2,1)) )
R = U@Vt
dets = np.linalg.det(R)
Vt[:,self.n_dims-1,:] *= dets[:,None]
R = U@Vt
return R
def _build_rhs( self, rotations: np.ndarray ):
R = (np.take( rotations, self._nbrs, axis=0 )+rotations[:,None])*0.5
tru_hood = (self._pnts.T[...,None]-np.take( self._pnts, self._nbrs, axis=1 ).transpose((1,0,2)))*self._wgts[:,None,:]
rhs = np.sum( (R@tru_hood.transpose((0,2,1))[...,None]).squeeze(), axis=1 )
return rhs
def _build_constraint_rhs( self, anchors: Dict[int,Tuple[float,float,float]] ):
f = np.zeros((self.n_pnts,self.n_dims),dtype=float)
f[self._anchors,:] = np.take( self._pnts, self._anchors, axis=1 ).T
for i,v in anchors.items():
if i not in self._anchors:
raise ValueError('Supplied anchor was not included in list provided at construction!')
f[i,:] = v
return f
The indexing was tricky to figure out. Error handling and input checking can obviously be improved. Something I'm not going into are the handling of anchor vertices and the overall quadratic forms that are being solved.
So does it Work?
Yes. Here is a 2D equivalent to the bar example from the paper:
import time
import numpy as np
import matplotlib.pyplot as plt
import arap
def grid_mesh_2d( nx, ny, h ):
x,y = np.meshgrid( np.linspace(0.0,(nx-1)*h,nx), np.linspace(0.0,(ny-1)*h,ny))
idx = np.arange(nx*ny,dtype=int).reshape((ny,nx))
quads = np.column_stack(( idx[:-1,:-1].flat, idx[1:,:-1].flat, idx[1:,1:].flat, idx[:-1,1:].flat ))
tris = np.vstack((quads[:,(0,1,2)],quads[:,(0,2,3)]))
return np.row_stack((x.flat,y.flat)), tris, idx
nx,ny,h = (21,5,0.1)
pnts, tris, ix = grid_mesh_2d( nx,ny,h )
anchors = {}
for i in range(ny):
anchors[ix[i,0]] = ( 0.0, i*h)
anchors[ix[i,1]] = ( h, i*h)
anchors[ix[i,nx-2]] = (h*nx*0.5-h, h*nx*0.5+i*h)
anchors[ix[i,nx-1]] = ( h*nx*0.5, h*nx*0.5+i*h)
deformer = arap.ARAP( pnts, tris, anchors.keys(), anchor_weight=1000 )
start = time.time()
def_pnts = deformer( anchors, num_iters=2 )
end = time.time()
plt.triplot( pnts[0], pnts[1], tris, 'k-', label='original' )
plt.triplot( def_pnts[0], def_pnts[1], tris, 'r-', label='deformed' )
plt.legend()
plt.title('deformed in {:0.2f}ms'.format((end-start)*1000.0))
plt.show()
The result of this script is the following:
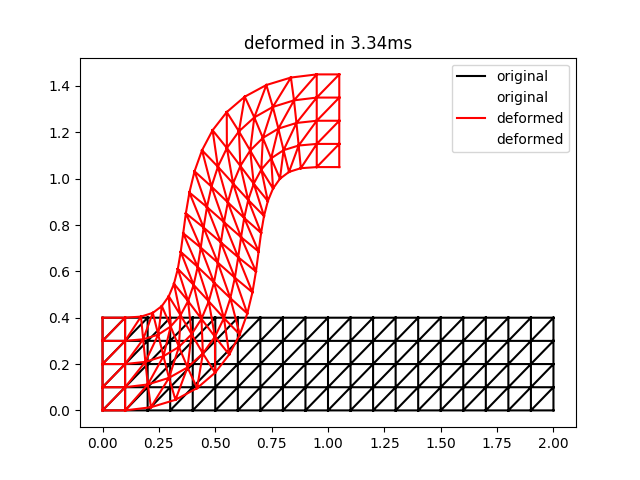
This is qualitatively quite close to the result for two iterations in Figure 3 of the ARAP paper. Note that the timings may be a bit unreliable for such a small problem size.
A corresponding result for 3D deformations is here (note that this corrects an earlier bug in rotation computations for 3D):
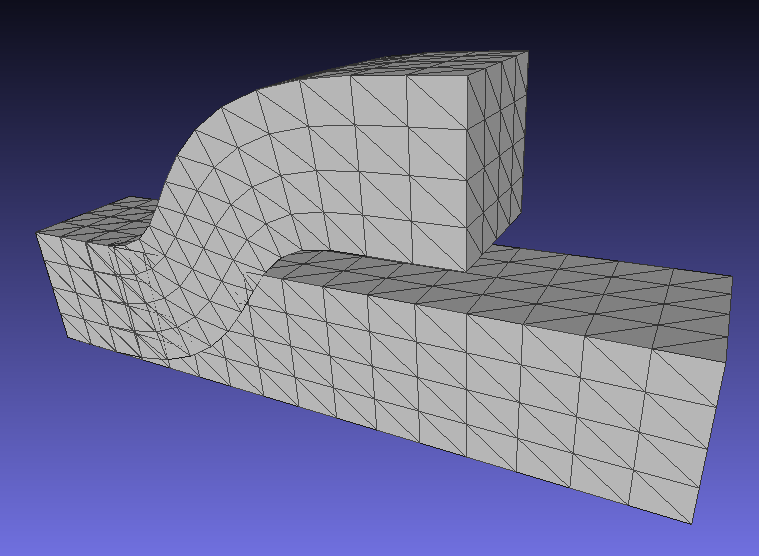

Nov 15, 2018
Here are a few operations for dealing with objective functions of matrix-valued variables:
Starting Definitions
Let \(A\) be a \(N\times M\) matrix with entry \(A_{ij}\) being the entry at row \(i\) and column \(j\). The \(i\)'th row of \(A\) is then \(A_{i*}\) and the \(j\)'th column is \(A_{*j}\).
Row and Column Vectorization
Define the column vectorization operator which returns a single column vector containing the columns of \(A\) stacked
\begin{equation*}
\newcommand{\cvec}{{\bf\mbox{cvec}}}
\cvec(A) = \begin{bmatrix}
A_{*1} \\
A_{*2} \\
\vdots \\
A_{*M}
\end{bmatrix}
\end{equation*}
likewise the row vectorization operator returns a single column vector containing the (transposes of) rows of \(A\) stacked:
\begin{equation*}
\newcommand{\rvec}{{\bf\mbox{rvec}}}
\rvec(A) = \cvec(A^T) = \begin{bmatrix}
A_{1*}^T \\
A_{2*}^T \\
\vdots \\
A_{N*}
\end{bmatrix}
\end{equation*}
In numpy, matrices are stored in row-major order, so the Python definition of these operaions is inverted as below:
def cvec( A ):
"""Stack columns of A into single column vector"""
return rvec(A.T)
def rvec( A ):
"""Stack rows of A into a single column vector"""
return A.ravel()
Similarly, the inverses of \(\cvec(A)\) and \(\rvec(A)\) unpack the vectorized values back into a matrix of the original shape:
\begin{align*}
\newcommand{\cmat}{{\bf\mbox{cmat}}}
\newcommand{\rmat}{{\bf\mbox{rmat}}}
\cmat\left( \cvec(A), M \right) = A \\
\rmat\left( \rvec(A), N \right) = A \\
\end{align*}
with corresponding python definitions:
def rmat( v, nr ):
"""Reshape vector v into matrix with nr rows"""
return v.reshape((nr,-1))
def cmat( v, nc ):
"""Reshape vector v into matrix with nc columns"""
return v.reshape((cols,-1)).T
Finally, define two additional operators. The first of these is the spread operator, which takes the Kronecker product between an identity matrix and the input matrix, resulting in copies of the input matrix along the diagonal:
\begin{equation*}
\newcommand{\spread}[2]{{\bf\mbox{spread}}_{#2}\left(#1\right)}
\spread{A}{r} = I_{r\times r} \otimes A = \begin{bmatrix}
A & 0 & \dots & 0 \\
0 & A & \dots & 0 \\
\vdots & \vdots & \ddots & \vdots \\
0 & 0 & \dots & A
\end{bmatrix}
\end{equation*}
The second of these is the split operator which reverses the order of the arguments to the Kronecker product, basically replacing each entry \(A_{ij}\) with \(A_{i,j} I_{r\times r}\):
\begin{equation*}
\newcommand{\split}[2]{{\bf\mbox{split}}_{#2}\left(#1\right)}
\split{A}{r} = A \otimes I_{r\times r} = \begin{bmatrix}
A_{11} I_{r\times r} & A_{12} I_{r\times r} & \dots & A_{1M} I_{r\times r} \\
A_{21} I_{r\times r} & A_{22} I_{r\times r} & \dots & A_{2M} I_{r\times r} \\
\vdots & \vdots & \ddots & \vdots \\
A_{N1} I_{r\times r} & A_{N2} I_{r\times r} & \dots & A_{NM} I_{r\times r}
\end{bmatrix}
\end{equation*}
Python implementation of these are one-liners that just call the existing numpy Kronecker product:
def spread( M, n ):
"""Make n copies of M along the matrix diagonal"""
return np.kron( np.eye(n), M )
def split( M, n ):
"""Replace each entry of M with a n-by-n identity matrix scaled by the entry value"""
return np.kron( M, np.eye(n) )
With these, it is possible to define matrix products. There are also some fun properties that come from the Kronecker product:
\begin{align*}
\spread{M}{n}^T &=& \spread{M^T}{n} \\
\spread{M}{n}^{-1} &=& \spread{M^{-1}}{n} \\
\split{M}{n}^T &=& \split{M^T}{n} \\
\split{M}{n}^{-1} &=& \split{M^{-1}}{n}
\end{align*}
Matrix Products
Let \(X\) be a \(N\times K\) matrix and \(Y\) be a \(K\times M\) matrix, then their product \(W = XY\) will be a \(N\times M\) matrix and the following are true:
\begin{align*}
\rvec(W) &=& \split{X}{M} \rvec(Y) \\
\cvec(W) &=& \spread{X}{M} \cvec(Y) \\
\rvec(W) &=& \spread{Y^T}{N} \rvec(X) \\
\cvec(W) &=& \split{Y^T}{N} \cvec(X)
\end{align*}
What these let you do is unbury any (single) term that's in the middle of some horrendous expression. For example, say there's an expression \(W = AXB\) with \(A \in \mathcal{R}^{N\times R}\), \(X \in \mathcal{R}^{R\times S}\) and \(B \in \mathcal{R}^{S \times M}\) and you want to isolate \(X\) as a column vector. Then you have:
\begin{align*}
\cvec(W) &=& \spread{A}{M} \split{B^T}{R} \cvec(X) \\
&=& \left( I_{M\times M} \otimes A \right) \left( B^T \otimes I_{R\times R} \right) \cvec(X) \\
&=& \left( I_{M\times M} B^T \right) \otimes \left( A I_{R\times R} \right) \cvec(X) \\
&=& \left( B^T \otimes A \right) \cvec(X)
\end{align*}
The step in the third line is the Kronecker product identity:
\begin{equation*}
\left( A \otimes B \right) \left( C \otimes D \right ) = \left( A C \right) \otimes \left( B D \right)
\end{equation*}
The same steps can also be done if you want row-major storage of \(X\):
\begin{align*}
\rvec(W) &=& \split{A}{M} \spread{B^T}{R} \rvec(X) \\
&=& \left( A \otimes I_{M\times M} \right) \left( I_{R\times R} \otimes B^T \right) \rvec(X) \\
&=& \left( A \otimes B^T \right) \rvec(X)
\end{align*}
In these cases the dimensions work out correctly but you don't need to worry too much about them since the dimensions of \(X\) are constrained by the column could in \(A\) and the row count in \(B\). This only applies when operating with them symbolically of course.
Frobenius Norm Problems
With this information, it becomes pretty easy to deal with Frobenius norm problems like:
\begin{equation*}
X = \mbox{argmin} \frac{1}{2} \| A X B - Y \|_F^2
\end{equation*}
I have found these frustrating in the past since I invariably end up in indexing hell. But it becomes much easier with the identities above. Applying them first of all isolates the X term to the right-hand-side of the expression and also converts the Frobenius norm to an \(L_2\) norm. Making the substitutions \(\tilde{x} = \cvec(X)\) and \(\tilde{y} = \cvec(y)\),
\begin{align*}
\tilde{x}_* &=& \mbox{argmin} \frac{1}{2} \| \left( B^T \otimes A \right) \tilde{x} - \tilde{y} \|_2^2 \\
&=& \mbox{argmin} \frac{1}{2} \tilde{x}^T \left( B \otimes A^T \right) \left( B^T \otimes A \right) \tilde{x} - \tilde{x}^T \left( B \otimes A^T \right) \tilde{y} + \mbox{constants} \\
&=& \mbox{argmin} \frac{1}{2} \tilde{x}^T \left( B B^T \otimes A^T A \right) \tilde{x} - \tilde{x}^T \left( B \otimes A^T \right) \tilde{y}
\end{align*}
This now looks like a standard least-squares problem, albeit one with this Kronecker product mashed into it. Setting the gradient to zero gives the solution as:
\begin{align*}
\tilde{x}_* &=& \left( B B^T \otimes A^T A \right)^{-1} \left( B \otimes A^T \right ) \tilde{y} \\
&=& \left( (B B^T)^{-1} \otimes (A^T A)^{-1} \right) \left( B \otimes A^T \right) \tilde{y} \\
&=& \left( (B B^T)^{-1} B \otimes (A^T A)^{-1} A^T \right) \tilde{y}
\end{align*}
This result has a similar form to \(AXB = \left( B^T \otimes A\right)\cvec(X)\), which can be used to get back to the matrix form of the expression:
\begin{align*}
X &=& (A^T A)^{-1} A^T Y \left( (B B^T)^{-1} B\right)^T \\
&=& (A^T A)^{-1} A^T Y B^T (B B^T)^{-1}
\end{align*}
It's also possible to come at this from a different direction
\begin{align*}
X &=& \mbox{argmin} \frac{1}{2} \| A X B - Y \|_F^2 \\
&=& \mbox{argmin} \frac{1}{2} \mbox{Tr}\left( (AXB - Y)(AXB - Y)^T \right)
\end{align*}
The gradient of this is given in the Matrix Cookbook.
\begin{equation*}
\frac{\partial}{\partial X} \frac{1}{2} \mbox{Tr}\left( (AXB - Y)(AXB - Y)^T \right) = A^T \left( A X B - Y \right) B^T
\end{equation*}
Setting the gradient of this to zero and isolating \(X\) gives the following:
\begin{align*}
A^T A X B B^T - A^T Y B^T = 0 \\
X = (A^T A)^{-1} A^T Y B^T (B B^T)^{-1}
\end{align*}
which matches the previous result.
Nov 02, 2018
This page describes a basic approach to extracting Euler angles from rotation matrices. This follows what OpenCV does, (which is where I got the process) but I also wanted to understand why this approach was used.
Here I'm considering rotation performed first around x-axis (\(\alpha\)), then y-axis (\(\beta\)) and finally z-axis (\(\gamma\)). For this, the rotation matrix is shown below. Other rotation orders are analogous.
\begin{equation*}
R(\alpha,\beta,\gamma) = \left[\begin{matrix}\cos{\left (\beta \right )} \cos{\left (\gamma \right )} & \sin{\left (\alpha \right )} \sin{\left (\beta \right )} \cos{\left (\gamma \right )} - \sin{\left (\gamma \right )} \cos{\left (\alpha \right )} & \sin{\left (\alpha \right )} \sin{\left (\gamma \right )} + \sin{\left (\beta \right )} \cos{\left (\alpha \right )} \cos{\left (\gamma \right )} & 0\\\sin{\left (\gamma \right )} \cos{\left (\beta \right )} & \sin{\left (\alpha\right )} \sin{\left (\beta \right )} \sin{\left (\gamma \right )} + \cos{\left (\alpha \right )} \cos{\left (\gamma \right )} & - \sin{\left (\alpha \right )} \cos{\left (\gamma \right )} + \sin{\left (\beta \right )} \sin{\left (\gamma \right )} \cos{\left (\alpha \right )} & 0\\- \sin{\left (\beta \right )} & \sin{\left (\alpha \right )} \cos{\left (\beta \right )} & \cos{\left (\alpha \right )} \cos{\left (\beta \right )} & 0\\0 & 0 & 0 & 1\end{matrix}\right]
\end{equation*}
To solve for the Euler angles, we can use a little bit of trigonometry. The best way, in my opinion, is to find pairs of entries that are each products of two factors. Using the identity \(\sin^2 + \cos^2 = 1\) can be used to isolate specific angles, e.g. \(R_{0,0}\) and \(R_{0,1}\) can be used to find \(\cos(\beta)\) which unlocks the rest of the angles:
\begin{align*}
(-1)^k \cos(\beta + k \pi) = \sqrt{ \cos^2(\beta) ( \sin^2(\gamma) + \cos^2(\gamma)) } = \sqrt{ R_{0,0}^2 + R_{1,0}^2 } \\
\cos(\beta) \approx \sqrt{ R_{0,0}^2 + R_{1,0}^2 }
\end{align*}
Pay attention to the first line: For any true value of \(\beta\), this formula will return the same value for \(\beta + k\pi\) where \(k\) is an arbitrary integer. This will in turn determine the angles \(\alpha\) and \(\gamma\) in order to be compatible. The approximate equality will return \(\beta \in [-\pi/2, \pi/2]\).
Once \(\cos(\beta)\) is found we can find the angles directly using atan2:
\begin{align*}
\beta = \mbox{atan2}( -R_{2,0}, \cos(\beta) ) \\
\alpha = \mbox{atan2}\left( \frac{R_{2,1}}{\cos(\beta)}, \frac{R_{2,2}}{\cos(\beta)} \right) \\
\gamma = \mbox{atan2}\left( \frac{R_{1,0}}{\cos(\beta)}, \frac{R_{0,0}}{\cos(\beta)} \right)
\end{align*}
The other issue occurs when \(|\cos(\beta)| = 0\), which causes division by zero in the equations for \(\alpha,\gamma\) (\(\beta\) is still well defined). In this case there are a number of options. What OpenCV does is to arbitrarily decide that \(\gamma = 0\), which means that \(\sin(\gamma) = 0\) and \(\cos(\gamma) = 1\). The formulas in this case are:
\begin{align*}
\beta = \mbox{atan2}( -R_{2,0}, \cos(\beta) ) \\
\alpha = \mbox{atan2}( -R_{1,2}, R_{1,1} ) \\
\gamma = 0
\end{align*}
There are, of course, other options. Python code implementing these operations is below, along with unit testing for the degenerate and non-degerate cases.
import unittest
import numpy
def x_rotation( theta ):
"""Generate a 4x4 homogeneous rotation matrix about x-axis"""
c = numpy.cos(theta)
s = numpy.sin(theta)
return numpy.array([
[ 1, 0, 0, 0],
[ 0, c,-s, 0],
[ 0, s, c, 0],
[ 0, 0, 0, 1]
])
def y_rotation( theta ):
"""Generate a 4x4 homogeneous rotation matrix about y-axis"""
c = numpy.cos(theta)
s = numpy.sin(theta)
return numpy.array([
[ c, 0, s, 0],
[ 0, 1, 0, 0],
[-s, 0, c, 0],
[ 0, 0, 0, 1]
])
def z_rotation( theta ):
"""Generate a 4x4 homogeneous rotation matrix about z-axis"""
c = numpy.cos(theta)
s = numpy.sin(theta)
return numpy.array([
[ c,-s, 0, 0],
[ s, c, 0, 0],
[ 0, 0, 1, 0],
[ 0, 0, 0, 1]
])
def xyz_rotation( angles ):
"""Generate 4x4 homogeneous rotation in x then y then z order"""
return numpy.dot( z_rotation(angles[2]), numpy.dot( y_rotation(angles[1]), x_rotation(angles[0]) ) )
def xyz_rotation_angles( R, eps=1e-8, check=False ):
"""Back out the Euler angles that would lead to the given matrix"""
if check and numpy.linalg.norm( numpy.dot( R.transpose(), R ) - numpy.eye(4) ) > eps:
raise ValueError('Input matrix is not a pure rotation, R\'R != I')
cos_beta = numpy.sqrt( R[0,0]**2 + R[1,0]**2 )
if numpy.abs(cos_beta) > eps:
alpha = numpy.arctan2( R[2,1]/cos_beta, R[2,2]/cos_beta )
beta = numpy.arctan2(-R[2,0], cos_beta )
gamma = numpy.arctan2( R[1,0]/cos_beta, R[0,0]/cos_beta )
return numpy.array((alpha,beta,gamma))
else:
alpha = numpy.arctan2(-R[1,2], R[1,1] )
beta = numpy.arctan2(-R[2,0], cos_beta )
gamma = 0
return numpy.array((alpha,beta,gamma))
class test_euler_angles(unittest.TestCase):
def test_angles(self):
"""Do fuzz testing on arbitrary rotations, limiting beta to valid range"""
for i in range(1000):
alpha = (numpy.random.rand()-0.5)*numpy.pi*2
beta = (numpy.random.rand()-0.5)*numpy.pi*0.9999
gamma = (numpy.random.rand()-0.5)*numpy.pi*2
ang = xyz_rotation_angles( xyz_rotation([alpha,beta,gamma]))
self.assertAlmostEqual( alpha, ang[0] )
self.assertAlmostEqual( beta, ang[1] )
self.assertAlmostEqual( gamma, ang[2] )
def test_degeneracies(self):
"""Do fuzz testing on the degenerate condition of beta = +- pi/2"""
for i in range(1000):
alpha = (numpy.random.rand()-0.5)*numpy.pi*2
beta = numpy.sign(numpy.random.randn())*numpy.pi/2
gamma = (numpy.random.rand()-0.5)*numpy.pi*2
R = xyz_rotation((alpha,beta,gamma))
ang = xyz_rotation_angles( R )
R2 = xyz_rotation( ang )
# check that beta is recovered, gamma is set to zero and
# the rotation matrices match to high precision
self.assertAlmostEqual( beta, ang[1] )
self.assertAlmostEqual( ang[2], 0 )
for j in range(16):
self.assertAlmostEqual( R.ravel()[j], R2.ravel()[j] )
if __name__ == '__main__':
unittest.main()
Nov 02, 2018
Starting with the \(4\times4\) homogeneous transform matrix with parameters \(\beta = [ \theta_x, \theta_y, \theta_z, t_x, t_y, t_z ]^T\) where rotations are performed in XYZ order and using the following substitutions:
\begin{align*}
c_x = \cos(\theta_x) \\
s_x = \sin(\theta_x) \\
c_y = \cos(\theta_y) \\
s_y = \sin(\theta_y) \\
c_z = \cos(\theta_z) \\
s_z = \sin(\theta_z) \\
\end{align*}
the vector function mapping a point \(p = [p_x, p_y, p_z, 1]^T\) in the body coordinate system to a point in the world coordinate system \(w = [w_x, w_y, w_z, 1]^T\) is:
\begin{equation*}
\begin{bmatrix} w_x \\ w_y \\ w_z \\ 1 \end{bmatrix} = F( p, \beta ) =
\left[\begin{matrix}c_{y} c_{z} & - c_{x} s_{z} + c_{z} s_{x} s_{y} & c_{x} c_{z} s_{y} + s_{x} s_{z} & t_{x}\\c_{y} s_{z} & c_{x} c_{z} + s_{x} s_{y} s_{z} & c_{x} s_{y} s_{z} - c_{z} s_{x} & t_{y}\\- s_{y} & c_{y} s_{x} & c_{x} c_{y} & t_{z}\\0 & 0 & 0 & 1\end{matrix}\right]\begin{bmatrix} p_x \\ p_y \\ p_z \\ 1 \end{bmatrix}
\end{equation*}
and the Jacobian with respect to the parameters \(\beta\) is:
\begin{equation*}
J_F = \left[\begin{matrix}p_{y} \left(c_{x} c_{z} s_{y} + s_{x} s_{z}\right) + p_{z} \left(c_{x} s_{z} - c_{z} s_{x} s_{y}\right) & c_{x} c_{y} c_{z} p_{z} + c_{y} c_{z} p_{y} s_{x} - c_{z}p_{x} s_{y} & - c_{y} p_{x} s_{z} + p_{y} \left(- c_{x} c_{z} - s_{x} s_{y} s_{z}\right) + p_{z} \left(- c_{x} s_{y} s_{z} + c_{z} s_{x}\right) & 1 & 0 & 0\\p_{y} \left(c_{x} s_{y} s_{z} - c_{z} s_{x}\right) + p_{z} \left(- c_{x} c_{z} - s_{x} s_{y} s_{z}\right) & c_{x} c_{y} p_{z} s_{z} + c_{y} p_{y} s_{x} s_{z} - p_{x} s_{y} s_{z} & c_{y} c_{z} p_{x} + p_{y} \left(- c_{x} s_{z} + c_{z} s_{x} s_{y}\right) + p_{z} \left(c_{x} c_{z} s_{y} + s_{x} s_{z}\right) & 0 & 1 & 0\\c_{x} c_{y} p_{y} - c_{y} p_{z} s_{x} & - c_{x} p_{z} s_{y} - c_{y} p_{x} -p_{y} s_{x} s_{y} & 0 & 0 & 0 & 1\\0 & 0 & 0 & 0 & 0 & 0\end{matrix}\right]
\end{equation*}
Python code for these respective operations is below:
def make_transform( beta ):
c_x = numpy.cos(beta[0])
s_x = numpy.sin(beta[0])
c_y = numpy.cos(beta[1])
s_y = numpy.sin(beta[1])
c_z = numpy.cos(beta[2])
s_z = numpy.sin(beta[2])
t_x = beta[3]
t_y = beta[4]
t_z = beta[5]
return numpy.array([
[c_y*c_z, -c_x*s_z + c_z*s_x*s_y, c_x*c_z*s_y + s_x*s_z, t_x],
[c_y*s_z, c_x*c_z + s_x*s_y*s_z, c_x*s_y*s_z - c_z*s_x, t_y],
[-s_y, c_y*s_x, c_x*c_y, t_z],
[0, 0, 0, 1]
])
def make_transform_jacobian( beta, p ):
c_x = numpy.cos(beta[0])
s_x = numpy.sin(beta[0])
c_y = numpy.cos(beta[1])
s_y = numpy.sin(beta[1])
c_z = numpy.cos(beta[2])
s_z = numpy.sin(beta[2])
t_x = beta[3]
t_y = beta[4]
t_z = beta[5]
p_x = p[0]
p_y = p[1]
p_z = p[2]
return numpy.array([
[p_y*(c_x*c_z*s_y + s_x*s_z) + p_z*(c_x*s_z - c_z*s_x*s_y), c_x*c_y*c_z*p_z + c_y*c_z*p_y*s_x - c_z*p_x*s_y, -c_y*p_x*s_z + p_y*(-c_x*c_z - s_x*s_y*s_z) + p_z*(-c_x*s_y*s_z + c_z*s_x), 1, 0, 0],
[p_y*(c_x*s_y*s_z - c_z*s_x) + p_z*(-c_x*c_z - s_x*s_y*s_z), c_x*c_y*p_z*s_z + c_y*p_y*s_x*s_z - p_x*s_y*s_z, c_y*c_z*p_x + p_y*(-c_x*s_z + c_z*s_x*s_y) + p_z*(c_x*c_z*s_y + s_x*s_z), 0, 1, 0],
[c_x*c_y*p_y - c_y*p_z*s_x, -c_x*p_z*s_y - c_y*p_x - p_y*s_x*s_y, 0, 0, 0, 1],
[0, 0, 0, 0, 0, 0]
])
I generated these using sympy to build the transformations and used a finite-difference Jacobian function to check the output.

